Experimental Man
Ultra self-tracking
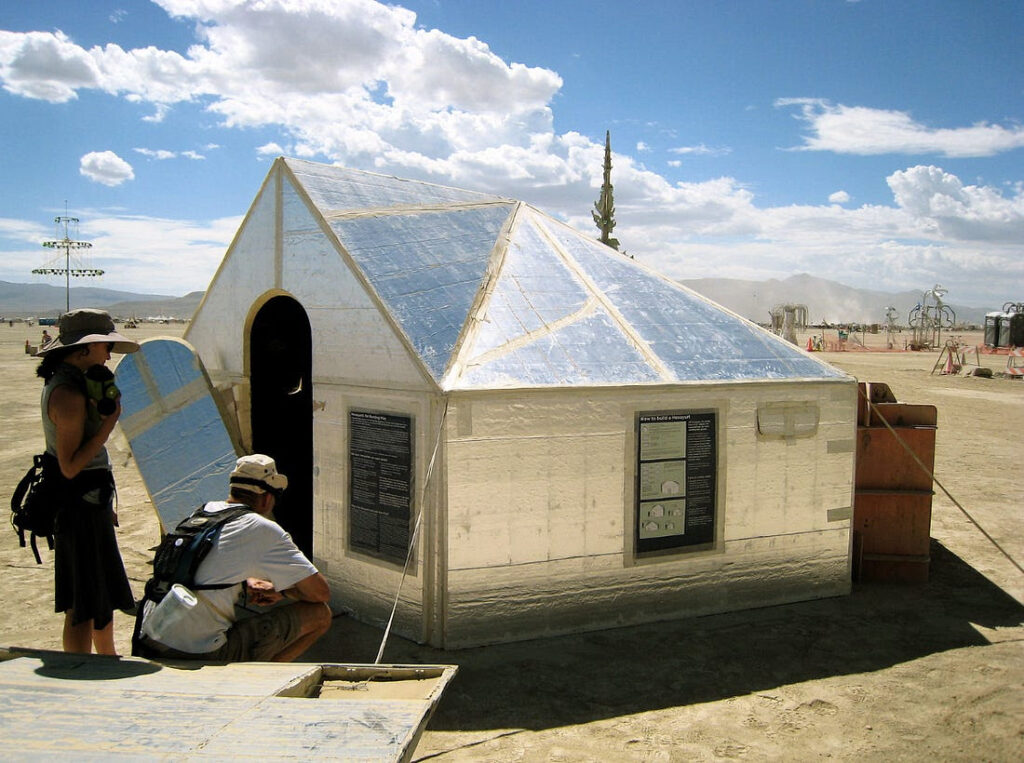
Someday you will be able to continuously measure your body in a hundred ways, and this constant data will transform your health. For the past few years David Duncan has been trying out this experiment. Although he is healthy, he’s subjected himself to every quantitative test he could find: multiple varieties of genome sequencing, measuring compounds in his blood, getting his brain scanned, tracking body pulses — and then he tried to correlate all this data. He calls himself the “experimental man.” The most fascinating part of his project was his attempt to measure the traces of environmental toxins left in his blood. I believe we will be following in his footsteps in the coming years. I started the site The Quantified Self just for this reason: in order to preview and discuss the tools for this kind of self-tracking (and I make a minor appearance in this book). Duncan’s account covers the plus and minus of this technology. He also gives us a clear sense of the potential for self-tracking and the immense difficulties we’ll have dealing with the data. I consider this book a very helpful and sobering glimpse of the future of health tools.
12/22/09Excerpt
Chris Austin believes that a much larger effort is needed, something akin to the Human Genome Project: perhaps the Human Envirogenomics Project? He and others believe that the only way to create meaningful envirogenetics data is through a large prospective cohort study, collecting DNA samples and information about exposure to a variety of environmental factors from five hundred thousand to a million participants who are followed for a number of years.
*
Hillis told me they took a picture of my proteome using the mass spectrometer--or, at least, a picture of the proteins swirling around in my blood on a certain day in April 2008, when I had my blood drawn. But it was a picture that included measurements at the atomic level of such complexity that it took about 24 gigabytes of storage space to hold all of the sample data (the picture I saw represents only about 1/24th of the total data from the sample)--that was fourteen hundred times the amount of digital space it took to store this entire book.
"It's a high-res picture of your whole proteome," said Ruderman.
We got up and walked over to an enormous flat-screen monitor on a wall of the lair, a TouchTable device invented by Hillis that models complex three-dimensional shapes on a flat screen--aircraft, buildings, cars, and proteomes. Ruderman clicked on my proteome file using his fingertips and pulled up functions that zoomed up and down the screen like an iPhone--although the touch-table technology had come first. Up popped a field of yellow dots that looked like a 3-D star field from outer space.
*
Meanwhile, huge chasms in our knowledge need to be filled before the Experimental Man will be complete down to every SNP, copy number variation, and synapse. Perhaps the biggest gap is the affect of the environment on our DNA, cells, organs, and bodies. Few of the tests I've taken for the Experimental Man project provide much useful information about how the environment interacts with genes, neurons, and proteomic systems and pathways. As cardiologist Eric Topol of Scripps told me, "You could almost say that giving genetic results without environmental data is inaccurate." The same is true about any system in the body, since the whole point of evolution has been to create defenses inside organisms to fend off the daily onslaught of the environment, from natural challenges such as UV rays and flu viruses to the thousands of toxic chemicals that we humans have unleashed into the air, the water, and the earth.
Experimental Man: What One Man's Body Reveals about His Future, Your Health, and Our Toxic World David Ewing Duncan 2009, 384 pages $18